This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Specific Aims Conclusions:
Aim #1: To determine the difference in FLNB between species with and without bones.
|
By seeing FLNB protein's phylogeny, I noticed two distinct groups: vertebrates and non-vertebrates. From this I proposed that FLNB could have another function in non-vertebrates. Though I did note that zebrafish were in the non-vertebrate group even though they are boney fish. To quickly see why zebrafish were in this group, I looked to see if there was any research done on FLNB in zebrafish. I found a lack of studies performed. So I did a STRING protein interaction network on zebrafish FLNB (Figure 1) and found that almost all interacting proteins are unknown. A lack of information on FLNB in zebrafish could explain why they are shown in the non-vertebrate group .
|
To determine why there are two distinct groups and if FLNB may play a different role in non-vertebrates, I performed a Clustal Omega alignment of the CH domain (Figure 2) and filamin domain (Figure 3) of FLNB to see if any differences are seen these two groups. The CH domain did not show any differences between the two groups and showed that there was much conservancy. But there was a difference between the two groups in the filamin domain. I noticed an amino acid change that was phosphorylated (Serine) in vertebrates and changed to an un-phosphorylated amino acid (Proline) in non-vertebrates. This could lead to great protein folding changes in most species, since there are several filamin repeats in most of these species. Because this shows that FLNB could have a different function, I propose doing further experiments on these non-vertebrate species. First I would propose tagging their specific FLNB protein with GFP to determine its localization in non-vertebrates and then go from there to determine the role of FLNB in these species.
Aim #2: To determine the role of FLNB in bone development with age.
From looking at the human FLNB protein interaction network (Figure 1: left) I was able to determine two groups of proteins that stood out: ubiquitin and fibronectin receptors. After learning the functions of these two groups of proteins as a whole, I became curious as to how these proteins change over time. I got curious after understanding that cartilage is degraded on bone joints with age as well as ossifies into bone tissue (1). To determine how these two groups of interacting proteins change over time, I proposed to start by identifying a good model organism to look at these two groups of interacting proteins. I performed a STRING interaction network of the mouse flnb protein (Figure 4: right) because the mouse flnb gene is very similar to the human FLNB gene and is a good vertebrate model organism. STRING showed that the mouse had all of the same proteins as the human in ubiquitin and fibronectin receptors with a few extra fibronectin receptors.
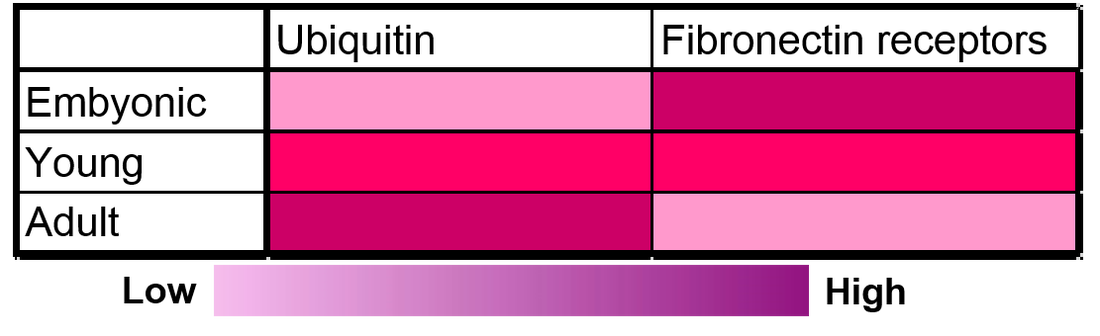
By finding the mouse to be a good model organism, I hypothesized how these interacting proteins with FLNB would change with age (Figure 5). I hypothesized that ubiquitin proteins would increase with age because we know that less cartilage is present in bone joints as we age (1). I hypothesized that fibronectin receptors would decrease with age because fibronectin receptors are involved in cartilage development. With less cartilage present as we age, I think that there would be less cartilage cells being made and localized.
To test my hypothesis, I would propose to tap tag these proteins to sort these proteins out. These proteins would be taken from mice cartilage and bone marrow tissue at embryonic, young, and adult stages. These tap tagged proteins would then run through mass spectrometry to see if any variation in the amount of these proteins occur with age.
By finding the mouse to be a good model organism, I hypothesized how these interacting proteins with FLNB would change with age (Figure 5). I hypothesized that ubiquitin proteins would increase with age because we know that less cartilage is present in bone joints as we age (1). I hypothesized that fibronectin receptors would decrease with age because fibronectin receptors are involved in cartilage development. With less cartilage present as we age, I think that there would be less cartilage cells being made and localized.
To test my hypothesis, I would propose to tap tag these proteins to sort these proteins out. These proteins would be taken from mice cartilage and bone marrow tissue at embryonic, young, and adult stages. These tap tagged proteins would then run through mass spectrometry to see if any variation in the amount of these proteins occur with age.
What are some future directions for Larsen Syndrome and FLNB?
After doing extensive research on Larsen Syndrome and FLNB, there were several future directions that came to mind that cover an array of areas. There was one area though that I believe would be an achievable, upcoming first goal for the medical and research field to achieve that would benefit medical staff and parents of those dealing with a patient or child with Larsen Syndrome. A great future direction, to me, would be to perform a large scale genomic study on those diagnosed with Larsen Syndrome to identify the most common variants along the FLNB gene. Past studies have only looked at small scale studies, with one looking at 8 patients and identifying 5 variants and the other looking at 20 patients and identifying 8 variants (2, 3). A large scale genome study could be done on 100 plus patients to get good results on resulting causative variants along the FLNB gene. The study will have to be done over a large area since Larsen Syndrome is a rare genetic disorder occurring in 1 in 100,000 people, for example in the tri-state area of the Midwest has a rough population number of 14 million people which means there could potentially be 140 patients with Larsen Syndrome in this area (though a little far fetched). In this study, all would be need from the patients would be a blood or tissue sample to run illumine Hi Seq 2000 genome sequencing on each of there respected FLNB genes. If a more supporting study was done that confirmed the most common variants to look for this could lead to possible prenatal screenings. If prenatal screenings could be done to diagnose Larsen Syndrome then tons of options could arise, options that I feel are very important to have. Options for the physicians by knowing prenatally if a baby is going to be born with Larsen Syndrome could help them take proper 3D ultra sounds to determine which surgeries might need to be done immediately after birth. Also physicians could plan for delivery be a cesarean section, to safely remove the baby to minimize further joint dislocations. Options for the parents with knowing that their baby will be born with Larsen Syndrome will allow them to talk with a genetic counselor to learn more about Larsen Syndrome and decide whether they want to terminate the pregnancy or continue. If the family decided to continue the pregnancy then they can better prepare for what's to come after birth and any surgeries that may be needed.
|
| ||||||||||||
References:
1.) "BCC: The skeleton, bones, and joints". Web. April 15, 2014.http://www.bbc.co.uk/schools/gcsebitesize/pe/appliedanatomy/2_anatomy_skeleton_rev4.shtml
2.) Bicknell, L. S., et. al., (2007). "A molecular and clinical study of Larsen Syndrome caused by mutations in FLNB". Journal of Medical Genetics, 44(1), doi:10.1136/jmg.2006.043687
3.) Krakow, D., et. al., (2004). "Mutations in the gene encoding filamin B disrupt vertebral segmentation, joint formation and skeletogenesis". Nature Genetics, 36(4), doi:10.1038/ng1319
2.) Bicknell, L. S., et. al., (2007). "A molecular and clinical study of Larsen Syndrome caused by mutations in FLNB". Journal of Medical Genetics, 44(1), doi:10.1136/jmg.2006.043687
3.) Krakow, D., et. al., (2004). "Mutations in the gene encoding filamin B disrupt vertebral segmentation, joint formation and skeletogenesis". Nature Genetics, 36(4), doi:10.1038/ng1319