This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What are protein motifs?
Protein motifs are sections of highly conserved regions that occur one or more times in an input protein sequence (1). Looking at the motifs of a protein can be useful in understanding which regions are important, such as is if a highly conserved region is mutated it could lead to a phenotypic effect. Motifs are also useful when comparing regions of a protein across other species. When looking for motifs of the FLNB protein, specifically in the CH and filamin domains, I used MEME. I used the Pfam database for finding the hidden markov model motif of these two domains as well, which can model entire alignments and is adept to representing amino acid insertions and deletions (2).
Protein motifs of FLNB: CH and filamin domains
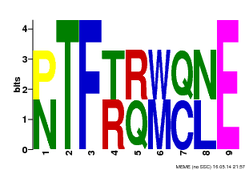
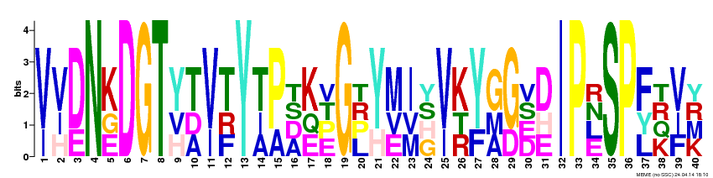
The FLNB protein CH and filamin domain motifs were generated by MEME, a useful database for identifying motifs in related sequences and compares the sequence over databases for human, dog, horse, as well as the mouse genome and motif sequences (1). To get these motifs on MEME, I inputted the FLNB CH domain and filamin domain protein FASTA sequences seperately with ">FLNB_HUMAN" for the first line of the sequence, as well as checked the "any number of repeats" option. The generated MEME motifs show the most conserved regions of the genomic sequence of these two domains (Figure 1 and Figure 3). The motifs for the protein are different than that of FLNB gene motifs for the motifs of the protein shown below use an amino acid code instead of the base pair code. Again the way to interpret these motifs is that the fewer the amino acids listed at a site the more conserved it is. For example in Figure 1, site 3 is very conserved for there is only one amino acid listed (Phenylalanine), compared to site 6 which is less conserved for two different amino acids can be found (Methionine and Tryptophan).
The hidden markov model motifs were generated by Pfam by pasting in the FLNB protein FASTA sequence under the "sequence search" tab. From these results, I clicked on the domain of my choosing which brought me to that domains information page. From there I found the hidden markov model motif by clicking on "HMM logo" on the left for the two domains (Figure 2 and Figure 4). The hidden markov model motifs are interpreted the same way as MEME motifs, but they have insertions and deletions shown on the motif as well as are much longer than the MEME motifs generated for the two domains of FLNB.
The hidden markov model motifs were generated by Pfam by pasting in the FLNB protein FASTA sequence under the "sequence search" tab. From these results, I clicked on the domain of my choosing which brought me to that domains information page. From there I found the hidden markov model motif by clicking on "HMM logo" on the left for the two domains (Figure 2 and Figure 4). The hidden markov model motifs are interpreted the same way as MEME motifs, but they have insertions and deletions shown on the motif as well as are much longer than the MEME motifs generated for the two domains of FLNB.
Analysis:
By submitting the MEME and Pfam FLNB CH and filamin protein sequences to generate motifs, I was able to visualize conserved regions of these domains from two different databases. This allowed me to see if there were any similarities or differences between the two types of databases. It is clearly seen that MEME generates shorter more conserved motifs of specific regions where as Pfam gives you a longer motif to visualize as well as insertions and deletions shown (2). After looking at these motifs from both domains, no specific amino acids stood out to me except a few phosphorylation amino acids that could be involved in protein folding. In the CH domain (Figure 1) there is a conserved Threonine (T) and in the filamin domain (Figure 3) there is a conserved Threonine (T) and Serine (S) amino acid in their motifs. If these regions were to be altered by a mutation it could lead to an abnormal protein via a change in protein folding.
References:
1.) "Current Protocols in Bioinformatics: Discovering Novel Sequence Motifs with MEME". Web. May 16, 2014. http://www.sdsc.edu/~tbailey/MEME-protocol-draft2/protocols.html
2.) "EBI: What are HMMs?". Web. February 27, 2014. http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/what-are-protein-signatures/signature-types/what-ar-1
2.) "EBI: What are HMMs?". Web. February 27, 2014. http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/what-are-protein-signatures/signature-types/what-ar-1