This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What are gene motifs?
|
Gene motifs are sections of highly conserved regions that occur one or more times in an input genomic sequence (1). Looking at the motifs of a gene can be useful in understanding which regions are important, such as is if a highly conserved region is mutated it could lead to a phenotypic effect. Motifs are also useful when comparing regions of a gene across other species. When looking for motifs of the FLNB gene I used MEME.
|
Motifs of the FLNB gene:
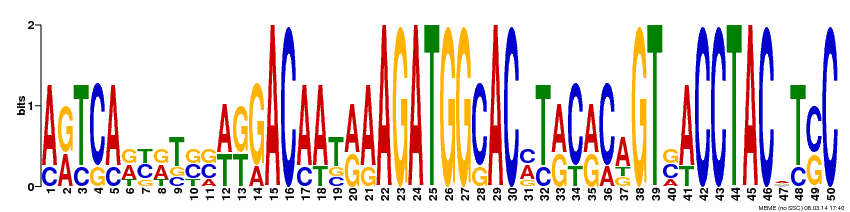
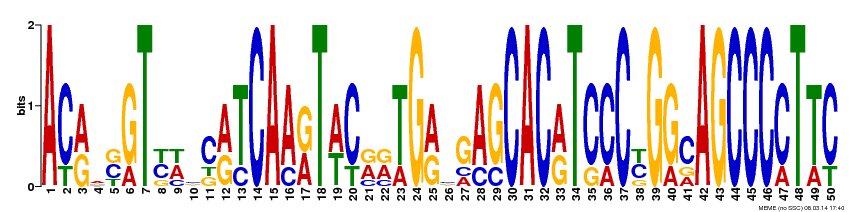
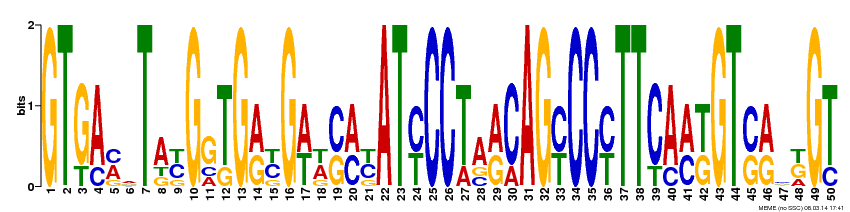
The FLNB gene motifs were generated by MEME, a useful database for identifying motifs in related sequences and compares the sequence over databases for human, dog, horse, as well as the mouse genome and motif sequences (1). To get these motifs on MEME, I inputted the FLNB gene cDNA FASTA sequence with ">FLNB_HUMAN" for the first line of the sequence, as well as checked the "any number of repeats" option. By doing this the 3 most conserved motifs resulted (Figure 2-4). Each base pair of the 50 base pair motifs are represented by a different color. The fewer the base pairs per area the more conserved that region is. For example in Figure 2, site 15 is very conserved and is found to always have the A (adenine) base pair. But at site 19 , in Figure 2, there are several possible base pairs identified at this region (C, G, T). I also submitted these motifs to GOMO a gene ontology site for motifs, but none of the motifs came up with any GOMO results.
Analysis of FLNB DNA motifs:
By submitting the FLNB cDNA FASTA sequence to MEME I was able to visualize regions of the FLNB gene that are highly conserved, and within these regions sites that are not so conserved. When looking at the three motifs, I saw that approximately ~37% of each of the motifs are highly conserved, where only one base pair occupies one site of the motif. Of these highly conserved base pair regions there are more C and G base pairs present than A and T. This is good supporting information for FLNB is G/C rich (2). This also goes along with one study that found that most mutations resulting in Larsen Syndrome is a missense mutation from a G to an A base pair in one of many identified mutation sites (2). Gene ontology the FLNB gene was completed and is found here since GOMO didn't assist any in identifying any gene ontology terms of the above motifs. This could have been because the motifs are only small sections of the FLNB gene, which is quite large.
References:
1.) "Current Protocols in Bioinformatics: Discovering Novel Sequence Motifs with MEME". Web. May 16, 2014. http://www.sdsc.edu/~tbailey/MEME-protocol-draft2/protocols.html
2.) Bicknell, L. S., et. al., (2007). "A molecular and clinical study of Larsen Syndrome caused by mutations in FLNB". Journal of Medical Genetics, 44(1), doi:10.1136/jmg.2006.043687
2.) Bicknell, L. S., et. al., (2007). "A molecular and clinical study of Larsen Syndrome caused by mutations in FLNB". Journal of Medical Genetics, 44(1), doi:10.1136/jmg.2006.043687