This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
What is phylogeny?
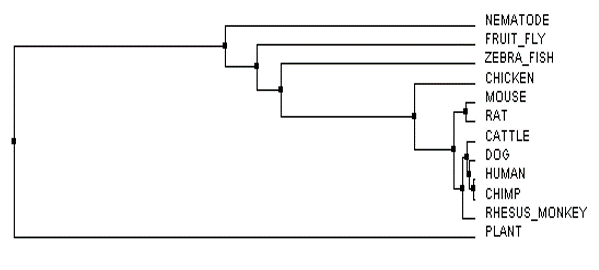
Phylogeny is the history of all species as they change through time (1). Phylogeny is typically expressed by a phylogenetic tree which shows how a group of species are related, whether they are closely related or have diverged from each other a long time ago. Species that share the same branch on a tree are more closely related than those that are on separate branches of the tree. For example in the tree shown in Figure 1, the mouse and the rat FLNB proteins are more closely related than the mouse and the chicken. The mouse and the rat do share a common ancestor with a chicken (this is noted by the node or the black dot) and you can see that it goes back in time a little to when this occurred in relation to the FLNB protein. You can see how all species on earth are all eventually connected together, for we all share one common ancestor.
What is the phylogeny of the FLNB protein?
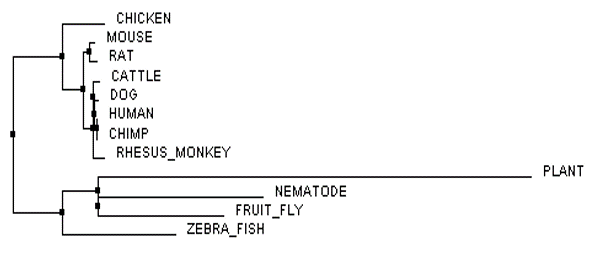
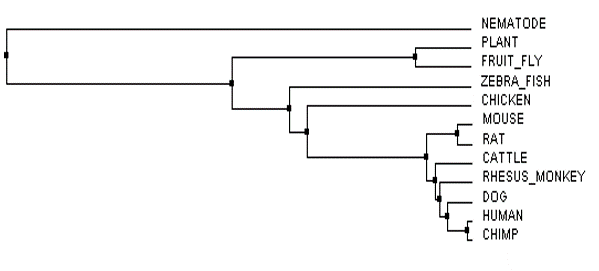
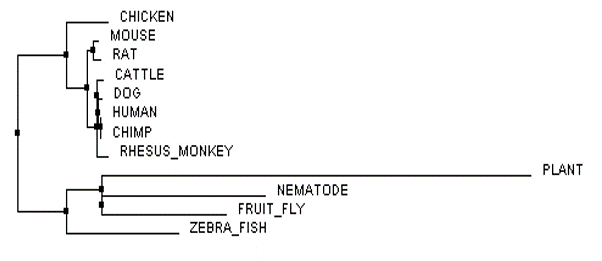
The phylogenetic trees of the FLNB protein were created by imputing a FASTA formatted text file (below) of all homologous FLNB protein sequences from the several species shown into Clustal Omega and Muscle. After the sequences were run through the databases, the phylogenetic tree was calculated to either Average Distance % Identity (Figure 3) or Neighbor Joining % Identity (Figure 4) as well as calculated with BLOSUM62 (Figure 1 and 2). The differences between Average Distance % Identity and Neighbor Joining % Identity is that Neighbor Joining % Identity analyzes the sequences to find a tree with the shortest branch lengths, where as Average Distance uses averages to create the clusters seen and then extends them to form a tree (2). The difference between Percent Identity and BLOSUM62 is that percentage identity uses a percentage between sequences at each aligned position by taking the "number of equivalent aligned non-gap symbols X 100 / smallest number of non-gap
positions in either of both sequences", where as BLOSUM62 matrix utilizes the sum of the BLOSUM62 score alignments for evolutionary divergent sequences. I attempted to create other phylogenetic trees using T-Coffee, Phylogeny.fr and TreeFam but my protein sequences were too large to do, the same as with my FLNB DNA sequences.
The phylogenetic trees tell us what species are most closely related by their specific protein sequence found, discussed under FLNB Protein Homology, and which are less related. Each of the trees also shows us which species the FLNB protein underwent a change first, second and so on through the evolution of time.
positions in either of both sequences", where as BLOSUM62 matrix utilizes the sum of the BLOSUM62 score alignments for evolutionary divergent sequences. I attempted to create other phylogenetic trees using T-Coffee, Phylogeny.fr and TreeFam but my protein sequences were too large to do, the same as with my FLNB DNA sequences.
The phylogenetic trees tell us what species are most closely related by their specific protein sequence found, discussed under FLNB Protein Homology, and which are less related. Each of the trees also shows us which species the FLNB protein underwent a change first, second and so on through the evolution of time.
Analysis of the FLNB protein phylogeny:
When it came to analyzing the phylogenetic trees of the FLNB protein one major thing stood out to me, and that is that there appears to be a split between the groupings of species with bones and species without bones. This is most easily seen in Figure 2 and Figure 4. More curiously, the zebrafish is a boney fish and is shown to be more closely related to the organisms without bones (plant, nematode, and fruit fly) than the other species with bones. This could pose that the FLNB protein could be involved in another role in organisms without bones, for what we do know is that FLNB is important in proper bone development and is a major protein binding actin filaments to the plasma membrane forming the actin cytoskeleton of cells. This poses something to look into further to see if research has been done in the species without bones to determine what the FLNB proteins role is in these species. It would also be interesting to try to determine what is going on with the FLNB protein in zebrafish that makes them more closely related to species that do not have bones when the zebrafish is clearly known as a boney fish. The FLNB protein could potentially have another role in zebrafish as well.
| flnb_protein_homology_fasta.pdf | |
| File Size: | 166 kb |
| File Type: | |
References:
1.) "Tree of Life Web Project: What is Phylogeny." Web. Feb, 20, 2014. http://tolweb.org/tree/learn/concepts/whatisphylogeny.html
2.) "Calculation of trees from alignments." Web. Feb, 20, 2014. http://www.jalview.org/help/html/calculations/tree.html
2.) "Calculation of trees from alignments." Web. Feb, 20, 2014. http://www.jalview.org/help/html/calculations/tree.html